Originally, they were designed for cell characterization, but they can be used in various applications, and more particularly in biomedical / medical image analysis.
1 - Size Zone Matrix (SZM)
2 - Multiple SZM
3 - Fuzzy SZM
3 - Distance (to border) Zone Matrix
4 - Orientation and Length Zone Matrix
Here is a Java (1.8) SOFTWARE to compute the features from these statistical matrices in both 2D and 3D. It can be used for radiomics, medical / biomedical images, etc.
How does it work?
First uncompress the file you have downloaded by clicking on the link above. You will find the file 'FiReTiTiLib.jar' and a directory 'lib'.
You can start the software by directly double clicking on the jar file FiReTiTiLib.jar or with the command line:
java -cp "lib/*.jar" -jar FiReTiTiLiB.jar
You will see appear a window that looks like that:
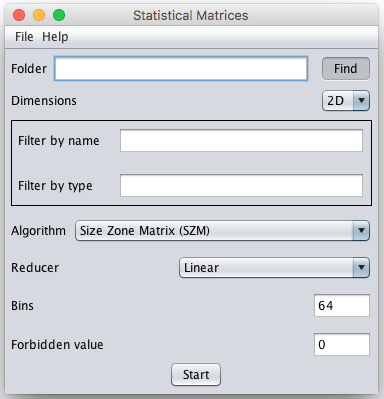
- Select the directory containing the images or the image sequences/stacks to process by using the button find, or copy/paste the link into the field "Folder".
- Select the data type: 2D for images, 3D for image sequences / stacks.
- (Optional) Use the fields "Filter by name" and/of "Filter by type" to process only the images/volumes that contains a specific name or extension/type.
- Select the algorithm (statistical matrix) to use.
- Select the gray level reducer.
- Enter the number of bins (gray level) for the regular reducers, or the fuzzyfication parameter for the fuzzy reducer.
- Enter a forbidden value. It's the value to ignore during the processing (matrix filling), generally the value representing the background, usually 0.
- Press "Start" and wait until a success (or failure, but let's hope not) message appears. It will indicate in which file the features were written.
- - The reducer "Fuzzy" is not compatible (yet) with all the algorithms.
- - The multiple versions as well as the fuzzy versions are much slower, particularly when using a high bins/fuzzy value.
- - If you start the same features extraction for two different files, do not forget to rename the first file, else it will be erased by the new one.
Do not hesitate to contact me if you need some explanations.